What happened
Here's a comprehensive summary of the MIT News article, "Using generative AI, researchers design compounds that can kill drug-resistant bacteria":
MIT Researchers Leverage Generative AI to Design Novel Antibiotics Against Drug-Resistant Bacteria
Publisher: MIT News
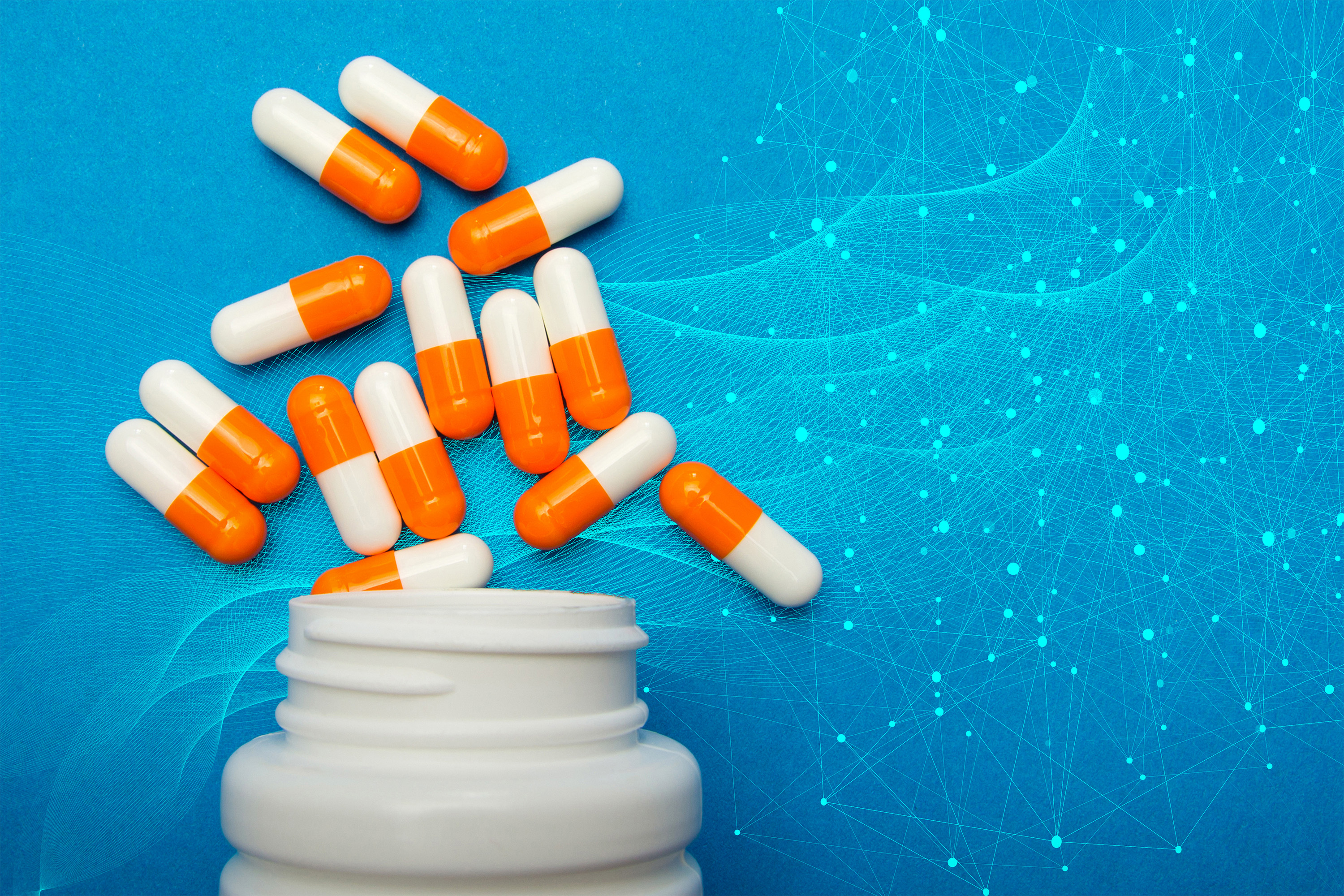
With help from artificial intelligence, MIT researchers have designed novel antibiotics that can combat two hard-to-treat infections: drug-resistant Neisseria gonorrhoeae and multi-drug-resistant Staphylococcus aureus (MRSA).Using generative AI algorithms, the research team designed more than 36 million possible compounds and computationally screened them for antimicrobial properties.
The top candidates they discovered are structurally distinct from any existing antibiotics, and they appear to work by novel mechanisms that disrupt bacterial cell membranes.This approach allowed the researchers to generate and evaluate theoretical compounds that have never been seen before — a strategy that they now hope to apply to identify and design compounds with activity against other species of bacteria.
“We’re excited about the new possibilities that this project opens up for antibiotics development. Our work shows the power of AI from a drug design standpoint, and enables us to exploit much larger chemical spaces that were previously inaccessible,” says James Collins, the Termeer Professor of Medical Engineering and Science in MIT’s Institute for Medical Engineering and Science (IMES) and Department of Biological Engineering.
Collins is the senior author of the study, which appears today in Cell. The paper’s lead authors are MIT postdoc Aarti Krishnan, former postdoc Melis Anahtar ’08, and Jacqueline Valeri PhD ’23.Exploring chemical spaceOver the past 45 years, a few dozen new antibiotics have been approved by the FDA, but most of these are variants of existing antibiotics.
At the same time, bacterial resistance to many of these drugs has been growing. Globally, it is estimated that drug-resistant bacterial infections cause nearly 5 million deaths per year.In hopes of finding new antibiotics to fight this growing problem, Collins and others at MIT’s Antibiotics-AI Project have harnessed the power of AI to screen huge libraries of existing chemical compounds.
This work has yielded several promising drug candidates, including halicin and abaucin.To build on that progress, Collins and his colleagues decided to expand their search into molecules that can’t be found in any chemical libraries. By using AI to generate hypothetically possible molecules that don’t exist or haven’t been discovered, they realized that it should be possible to explore a much greater diversity of potential drug compounds.
In their new study, the researchers employed two different approaches: First, they directed generative AI algorithms to design molecules based on a specific chemical fragment that showed antimicrobial activity, and second, they let the algorithms freely generate molecules, without having to include a specific fragment.
For the fragment-based approach, the researchers sought to identify molecules that could kill N. gonorrhoeae, a Gram-negative bacterium that causes gonorrhea. They began by assembling a library of about 45 million known chemical fragments, consisting of all possible combinations of 11 atoms of carbon, nitrogen, oxygen, fluorine, chlorine, and sulfur, along with fragments from Enamine’s REadily AccessibLe (REAL) space.
Then, they screened the library using machine-learning models that Collins’ lab has previously trained to predict antibacterial activity against N. gonorrhoeae. This resulted in nearly 4 million fragments. They narrowed down that pool by removing any fragments predicted to be cytotoxic to human cells, displayed chemical liabilities, and were known to be similar to existing antibiotics.
This left them with about 1 million candidates.“We wanted to get rid of anything that would look like an existing antibiotic, to help address the antimicrobial resistance crisis in a fundamentally different way. By venturing into underexplored areas of chemical space, our goal was to uncover novel mechanisms of action,” Krishnan says.
Through several rounds of additional experiments and computational analysis, the researchers identified a fragment they called F1 that appeared to have promising activity against N. gonorrhoeae. They used this fragment as the basis for generating additional compounds, using two different generative AI algorithms.
One of those algorithms, known as chemically reasonable mutations (CReM), works by starting with a particular molecule containing F1 and then generating new molecules by adding, replacing, or deleting atoms and chemical groups. The second algorithm, F-VAE (fragment-based variational autoencoder), takes a chemical fragment and builds it into a complete molecule.
It does so by learning patterns of how fragments are commonly modified, based on its pretraining on more than 1 million molecules from the ChEMBL database.Those two algorithms generated about 7 million candidates containing F1, which the researchers then computationally screened for activity against N.
gonorrhoeae. This screen yielded about 1,000 compounds, and the researchers selected 80 of those to see if they could be produced by chemical synthesis vendors. Only two of these could be synthesized, and one of them, named NG1, was very effective at killing N. gonorrhoeae in a lab dish and in a mouse model of drug-resistant gonorrhea infection.
Additional experiments revealed that NG1 interacts with a protein called LptA, a novel drug target involved in the synthesis of the bacterial outer membrane. It appears that the drug works by interfering with membrane synthesis, which is fatal to cells.Unconstrained designIn a second round of studies, the researchers explored the potential of using generative AI to freely design molecules, using Gram-positive bacteria, S.
aureus as their target.Again, the researchers used CReM and VAE to generate molecules, but this time with no constraints other than the general rules of how atoms can join to form chemically plausible molecules. Together, the models generated more than 29 million compounds. The researchers then applied the same filters that they did to the N.
gonorrhoeae candidates, but focusing on S. aureus, eventually narrowing the pool down to about 90 compounds.They were able to synthesize and test 22 of these molecules, and six of them showed strong antibacterial activity against multi-drug-resistant S. aureus grown in a lab dish. They also found that the top candidate, named DN1, was able to clear a methicillin-resistant S.
aureus (MRSA) skin infection in a mouse model. These molecules also appear to interfere with bacterial cell membranes, but with broader effects not limited to interaction with one specific protein.Phare Bio, a nonprofit that is also part of the Antibiotics-AI Project, is now working on further modifying NG1 and DN1 to make them suitable for additional testing.
“In a collaboration with Phare Bio, we are exploring analogs, as well as working on advancing the best candidates preclinically, through medicinal chemistry work,” Collins says. “We are also excited about applying the platforms that Aarti and the team have developed toward other bacterial pathogens of interest, notably Mycobacterium tuberculosis and Pseudomonas aeruginosa.
”The research was funded, in part, by the U.S. Defense Threat Reduction Agency, the National Institutes of Health, the Audacious Project, Flu Lab, the Sea Grape Foundation, Rosamund Zander and Hansjorg Wyss for the Wyss Foundation, and an anonymous donor.
Source coverage
Here's a comprehensive summary of the MIT News article, "Using generative AI, researchers design compounds that can kill drug-resistant bacteria":
MIT Researchers Leverage Generative AI to Design Novel Antibiotics Against Drug-Resistant Bacteria
Deeper analysis
Full source content
With help from artificial intelligence, MIT researchers have designed novel antibiotics that can combat two hard-to-treat infections: drug-resistant Neisseria gonorrhoeae and multi-drug-resistant Staphylococcus aureus (MRSA).Using generative AI algorithms, the research team designed more than 36 million possible compounds and computationally screened them for antimicrobial properties.
The top candidates they discovered are structurally distinct from any existing antibiotics, and they appear to work by novel mechanisms that disrupt bacterial cell membranes.This approach allowed the researchers to generate and evaluate theoretical compounds that have never been seen before — a strategy that they now hope to apply to identify and design compounds with activity against other species of bacteria.
“We’re excited about the new possibilities that this project opens up for antibiotics development. Our work shows the power of AI from a drug design standpoint, and enables us to exploit much larger chemical spaces that were previously inaccessible,” says James Collins, the Termeer Professor of Medical Engineering and Science in MIT’s Institute for Medical Engineering and Science (IMES) and Department of Biological Engineering.
Collins is the senior author of the study, which appears today in Cell. The paper’s lead authors are MIT postdoc Aarti Krishnan, former postdoc Melis Anahtar ’08, and Jacqueline Valeri PhD ’23.Exploring chemical spaceOver the past 45 years, a few dozen new antibiotics have been approved by the FDA, but most of these are variants of existing antibiotics.
At the same time, bacterial resistance to many of these drugs has been growing. Globally, it is estimated that drug-resistant bacterial infections cause nearly 5 million deaths per year.In hopes of finding new antibiotics to fight this growing problem, Collins and others at MIT’s Antibiotics-AI Project have harnessed the power of AI to screen huge libraries of existing chemical compounds.
This work has yielded several promising drug candidates, including halicin and abaucin.To build on that progress, Collins and his colleagues decided to expand their search into molecules that can’t be found in any chemical libraries. By using AI to generate hypothetically possible molecules that don’t exist or haven’t been discovered, they realized that it should be possible to explore a much greater diversity of potential drug compounds.
In their new study, the researchers employed two different approaches: First, they directed generative AI algorithms to design molecules based on a specific chemical fragment that showed antimicrobial activity, and second, they let the algorithms freely generate molecules, without having to include a specific fragment.
For the fragment-based approach, the researchers sought to identify molecules that could kill N. gonorrhoeae, a Gram-negative bacterium that causes gonorrhea. They began by assembling a library of about 45 million known chemical fragments, consisting of all possible combinations of 11 atoms of carbon, nitrogen, oxygen, fluorine, chlorine, and sulfur, along with fragments from Enamine’s REadily AccessibLe (REAL) space.
Then, they screened the library using machine-learning models that Collins’ lab has previously trained to predict antibacterial activity against N. gonorrhoeae. This resulted in nearly 4 million fragments. They narrowed down that pool by removing any fragments predicted to be cytotoxic to human cells, displayed chemical liabilities, and were known to be similar to existing antibiotics.
This left them with about 1 million candidates.“We wanted to get rid of anything that would look like an existing antibiotic, to help address the antimicrobial resistance crisis in a fundamentally different way. By venturing into underexplored areas of chemical space, our goal was to uncover novel mechanisms of action,” Krishnan says.
Through several rounds of additional experiments and computational analysis, the researchers identified a fragment they called F1 that appeared to have promising activity against N. gonorrhoeae. They used this fragment as the basis for generating additional compounds, using two different generative AI algorithms.
One of those algorithms, known as chemically reasonable mutations (CReM), works by starting with a particular molecule containing F1 and then generating new molecules by adding, replacing, or deleting atoms and chemical groups. The second algorithm, F-VAE (fragment-based variational autoencoder), takes a chemical fragment and builds it into a complete molecule.
It does so by learning patterns of how fragments are commonly modified, based on its pretraining on more than 1 million molecules from the ChEMBL database.Those two algorithms generated about 7 million candidates containing F1, which the researchers then computationally screened for activity against N.
gonorrhoeae. This screen yielded about 1,000 compounds, and the researchers selected 80 of those to see if they could be produced by chemical synthesis vendors. Only two of these could be synthesized, and one of them, named NG1, was very effective at killing N. gonorrhoeae in a lab dish and in a mouse model of drug-resistant gonorrhea infection.
Additional experiments revealed that NG1 interacts with a protein called LptA, a novel drug target involved in the synthesis of the bacterial outer membrane. It appears that the drug works by interfering with membrane synthesis, which is fatal to cells.Unconstrained designIn a second round of studies, the researchers explored the potential of using generative AI to freely design molecules, using Gram-positive bacteria, S.
aureus as their target.Again, the researchers used CReM and VAE to generate molecules, but this time with no constraints other than the general rules of how atoms can join to form chemically plausible molecules. Together, the models generated more than 29 million compounds. The researchers then applied the same filters that they did to the N.
gonorrhoeae candidates, but focusing on S. aureus, eventually narrowing the pool down to about 90 compounds.They were able to synthesize and test 22 of these molecules, and six of them showed strong antibacterial activity against multi-drug-resistant S. aureus grown in a lab dish. They also found that the top candidate, named DN1, was able to clear a methicillin-resistant S.
aureus (MRSA) skin infection in a mouse model. These molecules also appear to interfere with bacterial cell membranes, but with broader effects not limited to interaction with one specific protein.Phare Bio, a nonprofit that is also part of the Antibiotics-AI Project, is now working on further modifying NG1 and DN1 to make them suitable for additional testing.
“In a collaboration with Phare Bio, we are exploring analogs, as well as working on advancing the best candidates preclinically, through medicinal chemistry work,” Collins says. “We are also excited about applying the platforms that Aarti and the team have developed toward other bacterial pathogens of interest, notably Mycobacterium tuberculosis and Pseudomonas aeruginosa.
”The research was funded, in part, by the U.S. Defense Threat Reduction Agency, the National Institutes of Health, the Audacious Project, Flu Lab, the Sea Grape Foundation, Rosamund Zander and Hansjorg Wyss for the Wyss Foundation, and an anonymous donor.
How this page is built
Goose Pod turns cited reporting into a public episode summary first, then pairs that summary with audio playback so listeners can check the source material before they decide how deeply to engage.
The goal is to make this page useful as a news landing page first, while still giving listeners transcript access, related episodes, and direct links back to the original publishers.